Human Brain PHARMACOME
The Human Brain PHARMACOME project is pursuing a completely different strategy than the pharmaceutical industry in its search for an effective preventive therapy for Alzheimer's disease. Classical drug development typically starts from a single hypothesis for the development of the disease - and has been failing with this approach in the field of dementia research for years.
Our project, however, starts with a variety of disease mechanisms that are available as computable models and allow data and knowledge to be very quickly cross-referenced and linked to simulations. In a unique, holistic approach, we have generated the Human Brain PHARMACOME (HBP), a mechanistic model of all compounds active in the human brain, their targets (molecular targets), and their mechanisms of action. With the help of the HBP, already known and approved active substances, which were actually developed for completely different diseases, can be examined for their potential effectiveness in the human brain. Such a drug repurposing approach significantly shortens development times, since properties, efficacy profiles or toxicity have already been positively tested and approved. The HBP becomes a prediction machine for drugs whose activity spectrum suggests that they may be useful for the prevention of Alzheimer's disease.
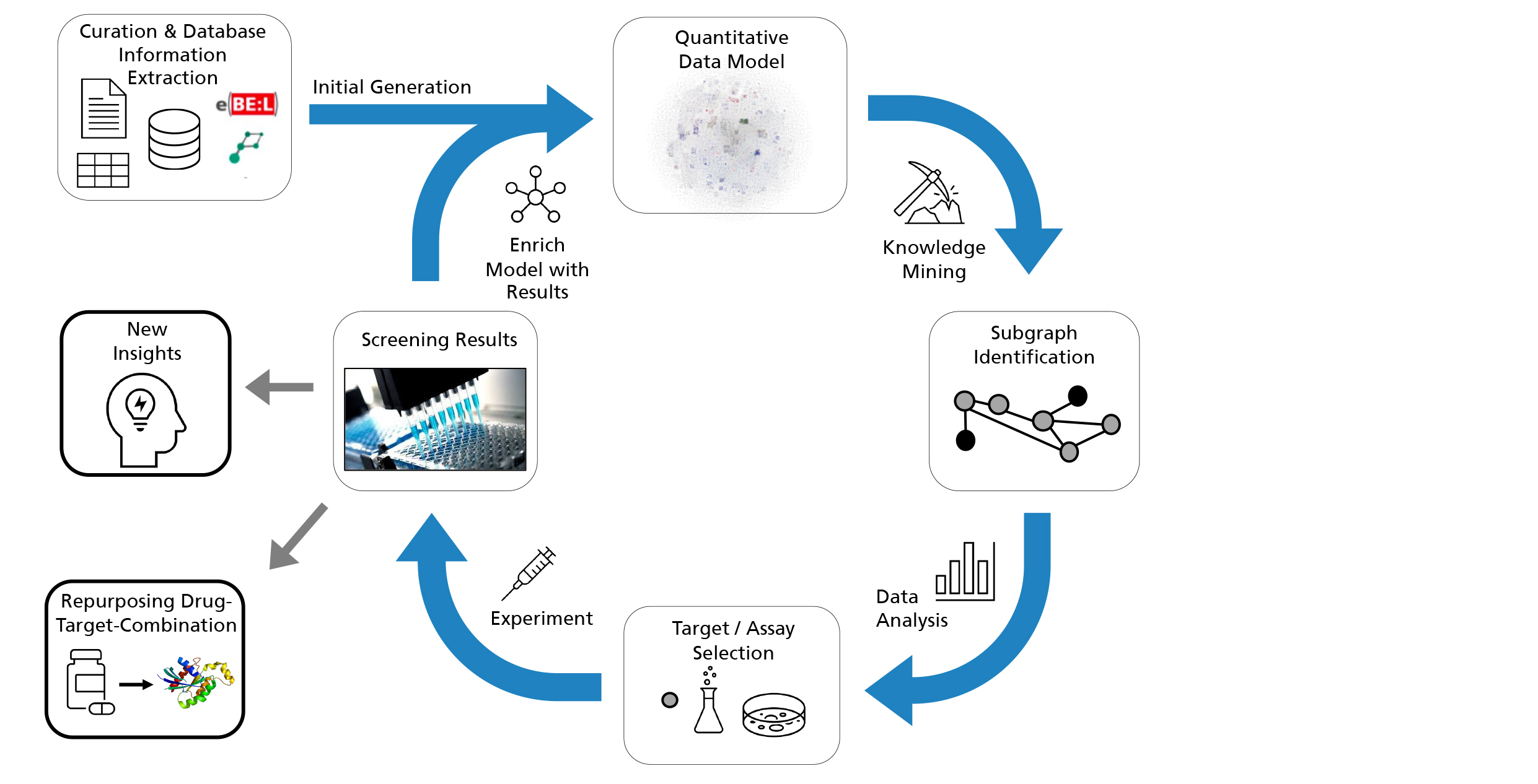
In the Human Brain PHARMACOME project, two computer science institutes are collaborating with a life science institute. With their information extraction technologies, the computer scientists create the prerequisite for a synopsis of the entire knowledge about active substances, their targets and their embedding in biological mechanisms. This is the HBP in its qualitative version. Using advanced information extraction techniques of numerical data from texts and tables, this qualitative model is extended by a quantitative component. Thus, we enable a smooth transition from knowledge-based, qualitative models to data-driven, quantitative models. Based on this quantitative version of the HBP, we perform mining procedures and simulations that enable us to predict and rank promising compounds. Candidate substances and their putative mechanism of action are derived from this in-silico prediction, which is based on intelligent methods. Colleagues at Fraunhofer IME in Hamburg, who have extensive experience in screening, i.e. systematically testing the efficacy of candidate substances, will then systematically test the suggestions in biochemical and cellular test systems known as assays. This unique combination of expertise from the fields of ICT and life science holds great potential, not least because of the unconventional, disruptive approach.
Our Use Case
We identified the histone deacetylase 6 (HDAC6) as a potential candidate for ptau modulation in AD pathological context. So far, it has been in the focus of cancer research (Seidel et al., 2015) because of its role in chromatin remodeling, cell motility and regulation of specific gene regulations (Aldana-Masangkay & Sakamoto, 2011). However, HDAC6 activity has also been shown to influence the structure of microtubules which makes it interesting in context of neurodegenerative diseases (d'Ydewalle et al., 2012). The relation identified from the HBP KB shows one new potential mechanism of action: HDAC6 shows a direct influence on acetylation of tau (Carlomagno et al., 2017). According to Carlomagno et al., HDAC6 mediated deacetylation of tau at certain points (like K321) favors phosphorylation of tau (Gorsky et al., 2016). Thus, an inhibition of HDAC6 might lead to a reduction of tau phosphorylation. Another study showed that HDAC6 activity at K280 and K281 positively correlated with the amount of tau protein (Cook et al., Hum Mol Genet, 2012). In both cases, the inhibition of HDAC6 should lead to a decrease in the harmful tau aggregation. However, there is also a report in which it has been shown that a direct acetylation of tau favors the generation of pathological tag aggregates (Cohen et al.,Nat Commun, 2011). In summary, it can be postulated that the dynamics of the post-translational modifications of tau, their mutual influence, and the downstream Effects on the formation of pathological "tau tangles" is not fully understood, especially in a humane context. It does appear to be a central molecule on which disease-relevant signal cascades in Alzheimer's disease are integrated.
Since the actual query language of the Human Brain PHARMACOME KB (Cypher) is very complex, simple Python notebooks were developed with which complex workflows can be mapped. These queries can be easily modified with regard to different questions, so that modulators of any molecular entities (contained in the HBP KB) and e.g. their post-translational modifications (PTM) can be queried in a similar way.
Publications
1. Mohammad Asif Emon: PS4DR: a multimodal workflow for identification and prioritization of drugs based on pathway signatures; 2019; BMC Bioinformatics; Preprint DOI: https://dx.doi.org/10.21203/rs.2.12791/v1
2. Özlem: GuiltyTargets: Target Prioritization with Network Representation Learning; 2019; Oxford Bioinformatics; DOI: https://doi.org/10.1101/521161
3. Charles Tapley Hoyt: Integration of Structured Biological Data Sources using Biological Expression Language; 2019; npj Systems Biology and Applications; bioRxiv; DOI: https://doi.org/10.1101/631812
4. Charles Tapley Hoyt: Re-curation and rational Enrichment of Knowledge Graphs in Biological Expression Language; 2019; Database, Volume 2019, 2019, baz068, https://doi.org/10.1093/database/baz068
Project managers
Prof. Dr. Martin Hofmann-Apitius, Fraunhofer SCAI,
Dr. Marc Jacobs (deputy)
Institutes involved
Fraunhofer SCAI, Fraunhofer IAIS, Fraunhofer ITMP
Project duration: 02/2018 – 06/2021